Root System Analyser - analysis of images and sequences of plant roots
We present a software package to analyse 2-dimensional images or image sequences of plant roots. The software measures root architectural traits such as root length, radius, connectivity, inter-root distances, or branching angles.
The following animations show the binary input images with the resulting predicted development of the root system. The predictions were performed by the software in a fully automatic way without any manual correction.
Artificial examples
The first animation shows an idealized (hand drawn) root system. The individual roots and their connectivity are recognized correctly. The animation is based on the assumption that each root grows at constant speed.
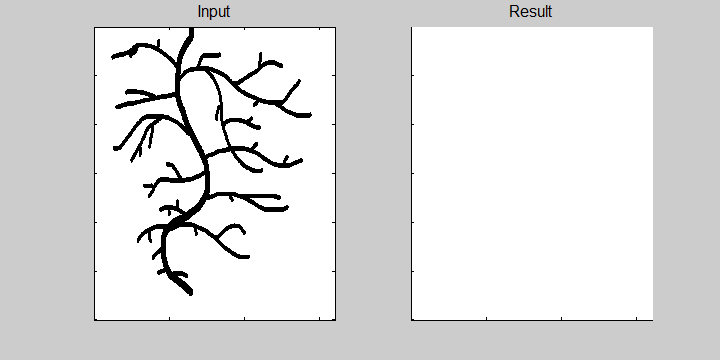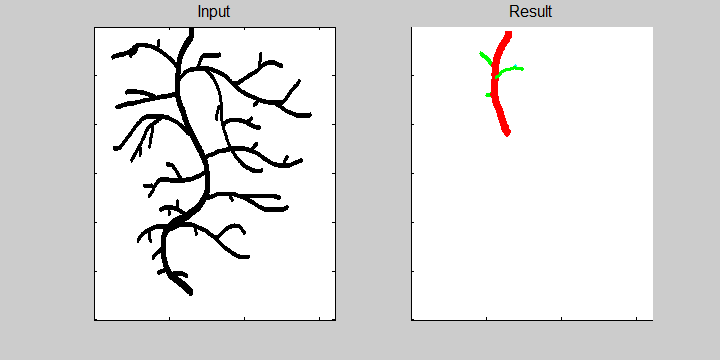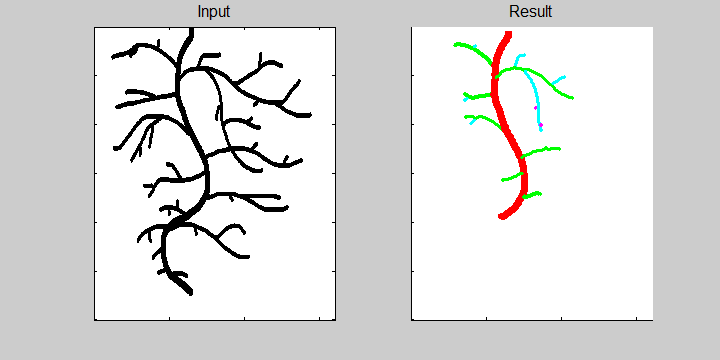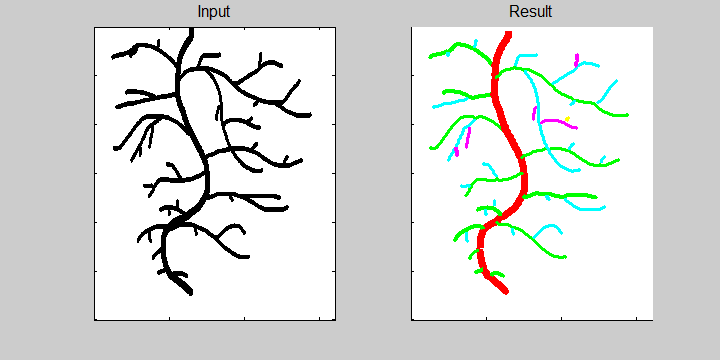
The following example demonstrates the ability of the software to recognize overlapping roots. This is essential for the analysis of a 2-dimensional image or image sequence.
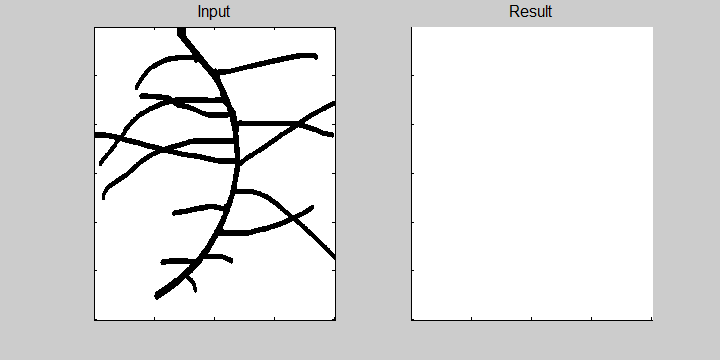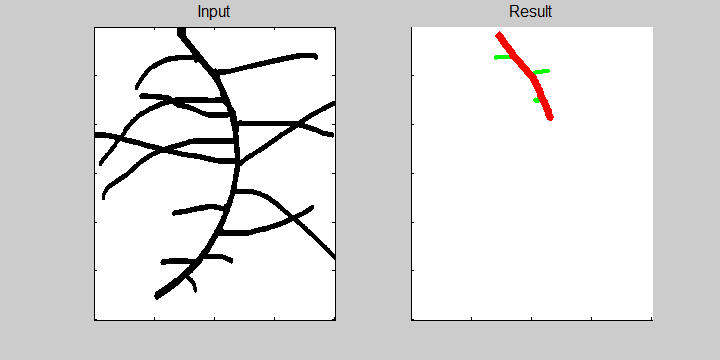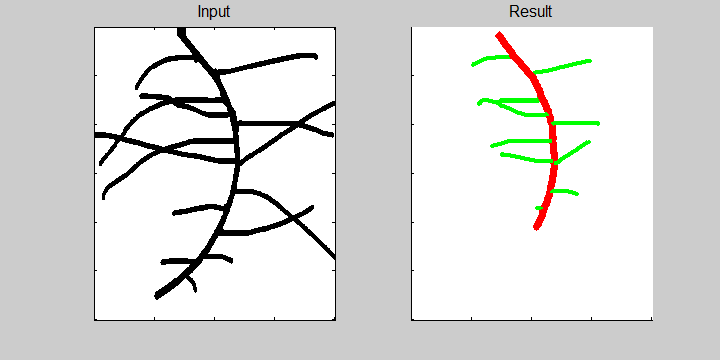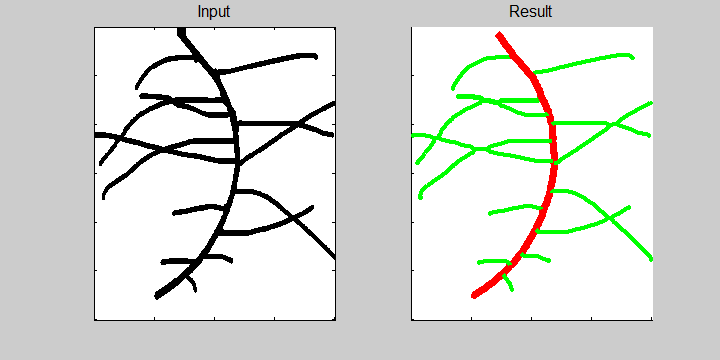
Images from neutron radiography
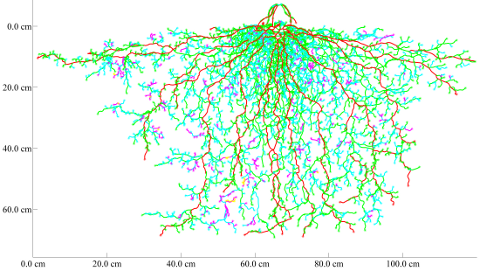
In the next animation the binary input is a single neutron radiographic image of a young lupine root system grown in soil in a thin container (27*26*15 cm). The majority of the roots (~95%) are recognized correctly. The animation shows the result without any correction by user interaction.

With neutron radiography it is possible to screen roots in situ in a non-destructive way. Therefore, it is possible to capture root development using an image sequence. Our root tracking software is capable to analyse roots in such image sequences and use this additional temporal information.
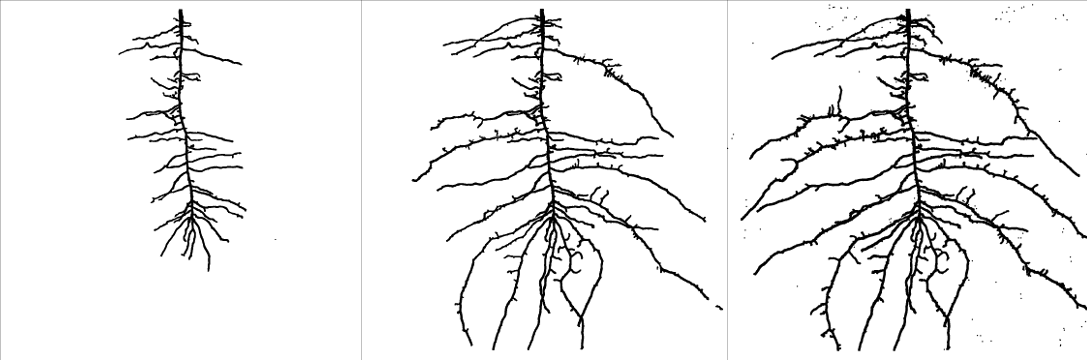
The above images show the development of the lupine plant after 12, 19, and 26 days of root growth. The predicted root architectural development is based on piecewise linear growth and is shown in this animation:
Fibrous root system
In the case of fibrous root architecture the user must supply the crown und brace roots manually. This is done by defining the base node and the tip node of each root.
How does it work?
- First we find a centreline of the binary image (skeletonization)
- We create a graph, where nodes are the root tips, branching point, or crossing points
- We start from the top node and 'fill' the graph by a growing root system, thus we assign each edge to one or more roots
- After this assignment we can compute root architectural parameters
More information
D Leitner, B Felderer, P Vontobel, and A Schnepf. Recovering root system traits using image analysis exemplified by two-dimensional neutron radiography images of lupine . Plant physiology, 164(1):24–35, 2014.Download
Version 2 (updated 19,10.2015) (including RSML support)- Standalone application (Windows)
- Documentation of Root System Analyser
- Matlab source code including GUI
- Matlab code including GUI
- Documentation of the GUI
- Matlab scripts for model parametrisation
Acknowledgments
The software was developed by Daniel Leitner and Andrea Schnepf. Daniel Leitner is recipient of an APART-fellowship of the Austrian Academy of Sciences at the Computational Science Center, University of Vienna. The work was supported by the Austrian Science Fund FWF (Grant Nos.: V220-N13).